| Crystal Structure Prediction and its Application in Earth and Materials Sciences |
The molecules we discussed so far are rigid or nearly rigid. Is it possible to use this approach to study the packings of flexible molecules? To prove this, we applied it to the prediction of crystal structure of butane-1,4-diammonium dibromide, in which Br and C
and C H
H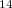 N
N
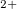 can be described as two molecular units that form the structure.
can be described as two molecular units that form the structure.
By using the experimental cell parameters 119, we indeed observed numerous structures with different conformations of the C H
H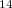 N
N
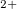 molecular ion. USPEX firstly found the energetically favorable conformation, and then identified the ground state structure at the 12th generation (about 500 structural relaxations): P2
molecular ion. USPEX firstly found the energetically favorable conformation, and then identified the ground state structure at the 12th generation (about 500 structural relaxations): P2 /c butane-1,4-diammonium dibromide. In this structure, as shown in Fig. 3.11, the organic hydrocarbon chains are found to pack in a herringbone-type stacking with hydrogen bonds to the Br
/c butane-1,4-diammonium dibromide. In this structure, as shown in Fig. 3.11, the organic hydrocarbon chains are found to pack in a herringbone-type stacking with hydrogen bonds to the Br . Each Br
. Each Br anion is surrounded by four –NH
anion is surrounded by four –NH
 groups. During the process of rotational mutation, both the orientation of the whole molecular group and its flexible torsional angles are allowed to change. A large fraction of rotation (
groups. During the process of rotational mutation, both the orientation of the whole molecular group and its flexible torsional angles are allowed to change. A large fraction of rotation ( 40%) of the molecules is found to greatly speed up the prediction. This success confirmed that our constrained evolutionary algorithm can be straightforwardly adapted to deal with flexible molecules.
40%) of the molecules is found to greatly speed up the prediction. This success confirmed that our constrained evolutionary algorithm can be straightforwardly adapted to deal with flexible molecules.
![\includegraphics[scale=0.4]{chapter3/pdf/Fig10.png}](images/img-0216.png)
 /c, Z=2). a), representation of network, view from a axis; b) Br
/c, Z=2). a), representation of network, view from a axis; b) Br coordination environment, view from b axis. For clarity, hydrogen atoms are not shown in (b). The C
coordination environment, view from b axis. For clarity, hydrogen atoms are not shown in (b). The C H
H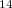 N
N
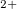 molecular ion has 6 flexible angles, and the unit cell of the stable polymorph contains 44 atoms.
molecular ion has 6 flexible angles, and the unit cell of the stable polymorph contains 44 atoms.