| Crystal Structure Prediction and its Application in Earth and Materials Sciences |
In the original version of USPEX 4, the stable crystal structure was assembled from individual atoms, which was also shown to work for atomic crystals and also for simple molecular systems (like carbon dioxide, water, urea). However, it is clear that for molecular crystals improvements of the efficiency can be made if the structure is assembled from whole molecules rather than individual atoms. This is confirmed by the present study. Our constrained global optimization method allows one to find the stable crystal structure of a given molecular compound, and provides a set of low-energy metastable structures at a highly affordable cost.
The reasons why evolutionary algorithms succeed in crystal structure prediction have been discussed before 23. As mentioned in Sec. II, in addition to these, the constrained global optimization fixes the molecular connectivity, and imposes the need for new variation operators (rotational mutation and softmutation), developed and described here.
For efficient and reliable polymorph prediction, the population of structures should be sufficiently diverse. A major difficulty in the prediction of molecular crystals is the large number of plausible candidate structures that can have very close free energies 122. Given the complexity of their energy landscape, high diversity of the population of the structures is mandatory for successful prediction of molecular crystal structures. The initial population is particularly important, and it is usually a good idea to add a number of random symmetrized structures in each generation, to keep sampling of the landscape diverse.
The presented algorithm provides not only the theoretical ground state, but also a number of low-energy metastable structures. With inclusion of zero-point energy and entropic contributions, such structures may become stable. Even if this does not happen, low-energy metastable structures have a relatively high chance to be synthesized at special conditions. While the DFT+D is today’s state of the art and its accuracy is often sufficient, for some systems (glycine), DFT+D is too crude, and more reliable approaches for computing the energy are needed. Under high pressure many of the difficulties disappear, because the vdW interactions (poorly accounted for by today’s ab initio methods) become relatively less important.
Clearly, the quality of the global minimum found by USPEX depends on the accuracy of the theory used for energy calculations and structure relaxation. Current levels of theory can be roughly divided into empirical, semi-empirical, and ab initio approaches. Accurate empirical force fields are appropriate for CSP, but reliable parameterizations are hard to generate for most molecules. In contrast to empirical force fields, ab initio calculations provide a more accurate and rigorous description without parameterization, but the calculations are much more time-consuming. In our prediction, we adopt the DFT+D level of theory, which combines "the best of both worlds", i.e. an accurate representation of intermolecular repulsions, hydrogen bonding, electrostatic interactions, and vdW dispersions. DFT+D proved to be reliable for most systems, but its results are not fully satisfactory for glycine. This shows that further improvements in theoretical calculations of intermolecular interactions energies are needed. In parallel with the improvement of methods for energy ranking, there is a need for efficient and reliable algorithms for global optimization of the theoretical energy landscape, and present work is an important development in this direction. In the present paper, we describe the most important ingredients of this method, and demonstrate how it enables affordable structure prediction for many complex organic and inorganic systems at ab initio level.
In summary, we have presented a new efficient and reliable approach for global energy optimization for molecular crystal structure prediction. It is based on the evolutionary algorithm USPEX extended to molecular crystals by additional variation operators and constraints of partially or completely fixed molecules. The high efficiency of this method enables fully quantum-mechanical structure prediction simulations to be performed at an affordable computational cost. Using this method, we succeeded in finding the stable structures for systems with various rigid molecular shapes (tetrahedral, linear, bent, planar and complex molecules), and chemical bonding (vdW bonding, ionic, covalent, metallic, weak and strong hydrogen bonding, 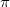 -
-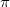 stacking, etc). We showed that even large systems can be efficiently dealt with by this approach, even at the ab initio level of theory. This new approach also has wide applicability to inorganic crystals containing clusters and complex ions.
stacking, etc). We showed that even large systems can be efficiently dealt with by this approach, even at the ab initio level of theory. This new approach also has wide applicability to inorganic crystals containing clusters and complex ions.